|
|
Hello flog!
For those of you who read all of my flog posts (I know there’s a solid number of you out there!) you’ve probably figured out by now that I love posting about numbers. So what’s today’s number?
Why, it’s 1948 of course!
Now this is the point that you might furtively look at wikipedia and say “I don’t understand what 1948 has to do with Echinacea. Everyone already knows that 1948 was a leap year starting on Thursday of the Gregorian calendar, the 1948th year of the Common Era (CE) and Anno Domini (AD) designations, the 948th year of the 2nd millennium, the 48th year of the 20th century, and the 9th year of the 1940s decade.” To which I would say that we are dealing with the number 1948, not the year.
No, 1948 is the number of seed packets of echinacea we x-rayed at the garden this week: and it’s only Wednesday! Through the combined efforts of many volunteers we are making some headway into the daunting task of figuring out which achenes have seeds in them and which do not. Look for updates soon about these number for our pollen limitation heads!
Michael
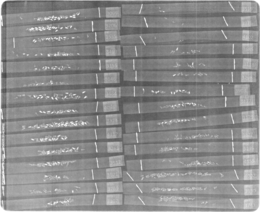
p.s., here’s a small sampler of what the xrays look like
 A look at our qGen_a xrays from 2013. There’s almost 900 images total in this folder (not nearly that many are shown here)
Hi all! Stuart and I are in the process of figuring out the best way to edit our x-ray images of Echinacea achenes so that it is easy to classify them:
- Are all achenes (full and empty) visible?
- Is it easy to tell between full and empty achenes?
- Can achenes within clumps be identified individually?
 Example x-ray image with achenes, before edited Here is an example of an x-ray image that is tricky to classify. All of these achenes are empty, so they are more translucent and harder to see. Also, many of these achenes are in clumps, so getting the correct count of empty achenes is a challenge.
 Example x-ray image before edits, with circled clump A single clump has been circled in green in this image. This clump has 3 empty achenes, but we want that to be more obvious. To edit our images, we have been working with the EBImage package in R. Here are a few examples.
 Edited example #1 This edit is helpful because it makes the background darker so that it’s easier to pick out achenes. The 3 in the clump are still kind of hard to pick out, though. Here is the code we used:
library(EBImage)
x <- readImage('https://echinaceaproject.org/wp-content/uploads/2018/04/x2mid.jpg')
display(x)
kern = 2000*makeBrush(99, shape = 'Gaussian', sigma = 5)
display(whiteTopHat(x, kern))

This example is blurred a bit, which is helpful for seeing achenes in their entirety. This is the code we used:
library(EBImage)
x <- readImage('https://echinaceaproject.org/wp-content/uploads/2018/04/x2mid.jpg')
display(x)
display(gblur(x, sigma=0.8))
There are endless possibilities if using multiple EBImage functions on one image, and we are looking for some guidance. If you have experience with EBImage, what do you think are the best combinations of functions for reaching our goals? Thanks for the help!
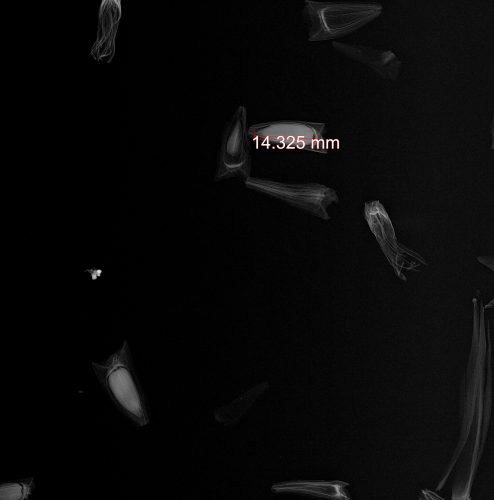
While looking for some information about the establishment of the Stipa experiment, I encountered an x-ray image of a few Stipa propagules (Hesperostipa spartea). Check it out…
 An x-ray image of a Stipa propagule (Hesperostipa spartea)
I also found some x-ray images of Echinacea angustifolia achenes. There are higher resolution images than the ones we now take for data analysis.
 An x-ray image of achenes & other stuff from an Echinacea angustifolia head with higher magnification
 An x-ray image of achenes & other stuff from an Echinacea angustifolia head  An x-ray image of achenes & other stuff from an Echinacea angustifolia head
In our lab we use UTHSCSA ImageTool to count thousands and thousands of achenes (fruits) every year on thousands of images. It has a straightforward interface and easily does exactly what we want: click cursor on an achene, a dot appears on the achene and the counter increments by one, click cursor on next achene, a new dot appears, counter increments, repeat until all achenes are tagged, then read the counter. Voila!
We are now going to classify achenes in x-ray images as full, partially full, or empty. We need software with the capability to tag three colors and make three separate counts per image. However, we don’t need software that is complicated, and can do everything. We want software that can do this one thing efficiently.
ImageJ will work, but it is clunky, taking many steps to set up the counting on each image. Can anyone recommend software suitable for our needs?
I found a few software packages to investigate, listed below, but would appreciate any advice or leads on other software.
http://fiji.sc/Welcome
http://marvinproject.sourceforge.net/en/index.html
http://itk.org/
http://imglib2.net/
http://imagej.net/ImageJ2
http://www.jmicrovision.com/
http://loci.wisc.edu/software/imagej
http://meesoft.logicnet.dk/Analyzer/
http://icy.bioimageanalysis.org/
Click to enlarge the below images.
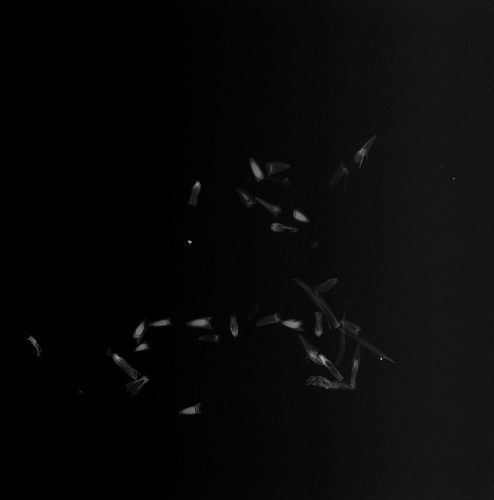
 X-ray image of Echinacea achenes
We count 12 full, 18 empty, and zero partially filled fruits (achenes).
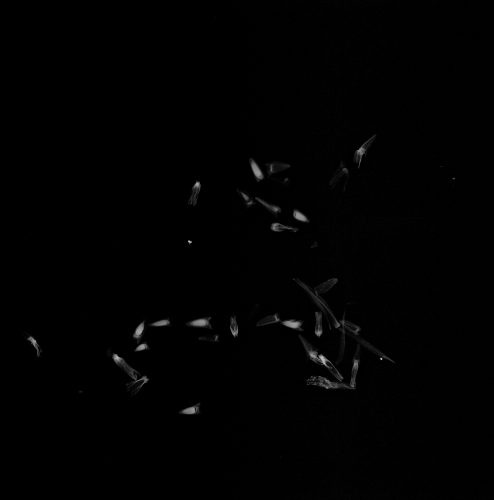
 Image of all achenes from Echinacea head GP-6008
How many achenes do you count?
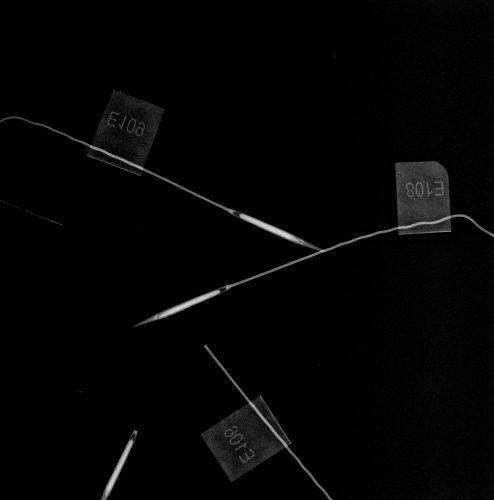
We took some high-quality images of Echinacea achenes for our q2 experiment this fall; an example is below. Notice how easy it is to distinguish empty achenes from those with embryos. By darkening the room and removing the opaque film, we were able to use lower levels of xrays for a shorter duration than we have previously. This plate was exposed to 12kV x-rays for 4s. We used long, thin glassine envelopes to facilitate counting. Notice also that the laser-printed labels reveal the packet IDs.

X-ray image of 30 packets of achenes from
Echinacea angustifolia. Click on thumbnail to enlarge.
What is a typical radiation dose experienced by an Echinacea seed when we x-ray Echinacea fruits to assess seed set?
We usually put the seeds on the bottom tray and the setting 10 s @ 18 kV. According to the documentation on dosage for our x-ray machine 18 kV outputs 292.6 R/h when the dosimeter probe is 57.2 cm away (that's the shelf with 1:1 magnification). The dose unit quote here R is Roetngen.
A little arithmetic can tell us total Roentgens for a ten second exposure:
( 10 s frac{1 h}{60 min} frac{1 min}{60 s} 292.6 frac{R}{h} )
In R code, that's
10 * 1/60 * 1/60 * 292.6
## [1] 0.8128
0.8128 R (Roentgen)
We may put the seeds on a higher shelf for more magnification, maybe 1:1.5 or the 1:2. We can enter dosages from each shelf from the documenation on exposure levels by shelf to estimate how much higher the dose is.
lvl <- c(1, 1.5, 2, 3, 4, 5)
RPM <- c(7.815, 20.55, 42.55, 115.45, 232, 397)
data.frame(shelf = lvl, dose = RPM)
## shelf dose
## 1 1.0 7.815
## 2 1.5 20.550
## 3 2.0 42.550
## 4 3.0 115.450
## 5 4.0 232.000
## 6 5.0 397.000
The dose on the 2x magnification shelf is RPM[3]/RPM[1] = 5.4447 times greater than the dose on the 1:1 shelf. Doubling the magnification generally should increase the dose by a factor of 5.4. Let's check: the dose on the 4x shelf is RPM[5]/RPM[3] = 5.4524 times greater than the dose on the 2x shelf. Also, the dose on the 3x shelf is RPM[4]/RPM[2] = 5.618 times greater than the dose on the 1.5x shelf. Close.
The expected dose at the stadard settings is 0.82 R on the bottom shelf and 0.8128 * 5.5 = 4.4704, or about 4.5 R on the 2x shelf.
Here is Sebastian Di Clemente’s final report on the main project of his internship:
|
|