In early December, NSF informed us that The Echinacea Project will be funded from April 2011 – April 2016!
|
||||
|
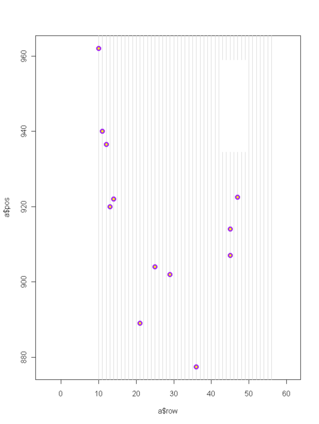
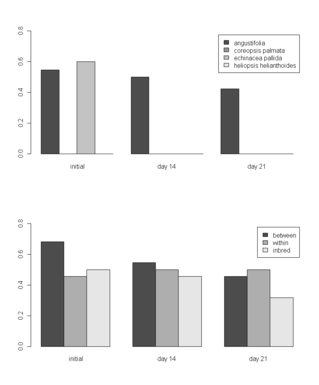
21 September 2010. Jennifer Ison defended her dissertation on Friday and gave a public seminar to a packed house at UIC on Tuesday. Congratulations! Her dissertation title is “Pollination of Echinacea angustifolia: effects of flowering phenology and spatial isolation.” Jennifer started with the Echinacea Project in 2003, right after graduating from St. Olaf College. She is heading to a post-doc position with Art Weis in Toronto, starting 1 October. Only about 13 heads left to harvest in the common garden experiment! I attached a pdf of listing all heads with no gbag info… HarvestList2010FinalHarvest.pdf Here’s a map of unharvested head locations in the main experiment (there’s one more head in the 99S garden). This R script, hillaryLookAtAphids.r, allows one to view graphs of growth of aphid clones in Lauren and Hillary’s experiment. Tag #1028 was just found in the ground near the SW corner of Hjelm house porch. Ian wants to quantify the overlap of flowering time between all pairs of plants in his experiment. This following R script reads his file with the flowering schedule of all 31 plants in his experiment and writes a file called ianPhenPairs.csv that has the flowering schedule of every possible pair combination (one per line). Note that there is a separate record for each plant as a sire and dam.
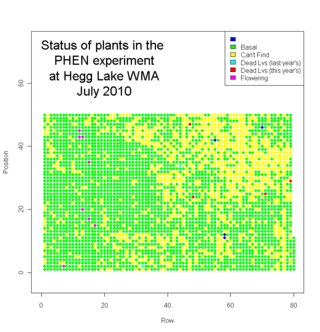
We finished measuring plants in Jennifer’s Phenology plot at Hegg Lake WMA this Friday. We measured many plants.
We measured about 2700 plants. Eight plants flowered this year. We visited another 1300 locations where we couldn’t find a plant (mostly because they had died). Here’s a map of the plot with the status of each location (click to see a bigger version)… In addition to Echinacea angustifolia, we saw some good prairie plants in the plot, including lead plant (Amorpha canescens), yellow lady-slipper (Cypripedium calceolus), prairie rose (Rosa arkansana), Missouri goldenrod (Solidago missouriensis), silver leaf scurf pea (Psoralea argophylla), and many others. It was a big job, but we were quite efficient. We laid out 50m tapes on every other row to help guide us. Four of us went out on Wednesday to flag positions 1 and 50 for all rows. That took ~2h. We started flagging positions 10, 20, but that was unnecessary. On Thursday we all went out and measured from 2-4 pm. On Friday we did two shifts: 10 -12 and 2:30 -5. We were happy to be done.
On the way out we removed a weed that we had noticed the day before–spotted knapweed. We were careful not to touch it because it can be a skin irritant. I’d never seen this plant in the study area before. Heading home.
What a great way to end the week. It was Katie and Laura’s last day. They are heading back to the Chicago Botanic Garden to prepare their posters and talks. This afternoon we are going to measure plants in Jennifer’s Phenology plot at Hegg Lake WMA. Here is a script that makes a datasheet that assigns us rows to measure. The order is approximately 1 – 80, but they are slightly mixed up (just to keep us on our toes): measurePHENatHeggLakeRows.r |
||||
|
© 2026 The Echinacea Project - All Rights Reserved - Log in Powered by WordPress & Atahualpa |
||||