|
|
Hello flog!
Unfortunately for all you T-Swift fans out there, I will not actually be devoting much of this blog to her… Instead, I am going to talk about the fascinating world of R, a statistical analysis coding program that allows you to organize and graph your data.
Since my time at the garden is coming to an end this week, I have begun to learn how to analyze my data within R. R can be a little complicated because often times even a Blank Space can mess up an entire line of code. However, with lots of help (like lots, from R genius Michael) I was able to organize and even plot some of my results! An important lesson- if your code doesn’t work the first time, Shake It Off and try again!
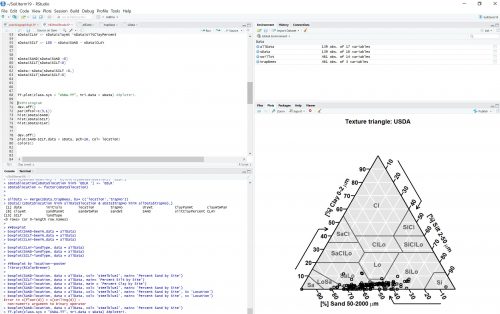
For my project, I am aiming to create two different graphs, a series of histograms and soil texture triangles. A histogram will show the amount of sand, silt, and clay within each of the 8 sites. Then to understand the texture differences, a soil triangle will plot the percentages of sand, silt, and clay in what type of soil they correlate to. For example, 45% sand, 40% silt, and 10% clay might fall at the bottom of the triangle in the sandy loam portion. Who knows maybe I’ll have Fifteen graphs by Friday!
 A preview of my graphs! Even though R can be a pretty Delicate program, it is extremely helpful! But being able to efficiently code has always been one of my Wildest Dreams. So, for now, there’s no Bad Blood between R and myself.
T-Swift out, till next time!
Hi Flog!
So, remember the micro- pipette method I mentioned a couple days ago? Well, as the name implies, everything is micro- the sample size, the vials, the pans, and the numbers. The process begins by weighing out 2 grams of each soil sample. Each sample is put in individual “tornado tubes” AKA vials filled with 20 ml of both water and sodium hexametaphosphate (try saying that ten times!). Vials are then shaken for approximately 30 minutes to create a uniform solution. Then, drum roll please…., comes the macro part of the process, the settling time. A 1 hour and 45 minute settling time… But this wait means I have time to write to all of you!
After the wait, comes the final part of the micro-pipette process that involves separating the clay and sand components of each sample into two different miniature pie pans. That means that 10 samples = 20 pans, 20 samples =40 pans, and 25 samples = 50 pans, etc etc etc… Guess how many samples I did today??!! 50! That’s 100 mini soil pies! Sometimes I feel like a professional baker making this many pies!


Anyway, as my first week at the garden comes to an end, I’ve developed more efficient methods and routines to help me be as productive as possible .I might even get close to finishing all 300 samples by the end of next week. I’ve also been introduced to an amazing community of scientists and can’t wait to learn more about their research!
Till next week!
Hey hey hey! My name is Anna and I was a summer intern on the Echinacea Project in high school. Now, as a college freshman, I am currently exploring another side of the project at the Chicago Botanic Garden lab! For my two weeks in Chicago, I will be working on the soil component of Kristen Manion’s project. Her study considers numerous elements of native bees’ habitat in hopes to determine their nesting preferences.
Over the summer, soil samples from 8 sites were collected and each site had three different land uses: old field, native prairie, or restoration. In total there are 328 samples to be analyzed through the micropipette soil texture method! The micropipette method involves separating the soil into its clay and sand components and then calculating the percentage of sand, silt, and clay for each 2-gram sample. By the end, I am aiming to create two soil triangles and answer these questions :
- Does the soil type differ between bee presence/absence?
- Does the soil type differ between land use types (remnant and prairies and old fields)?
My other goal is to simply gain experience in a lab by being exposed to a variety of techniques through Kristen’s and the Echinacea Project’s methods. However, I also have a couple of personal goals… like eating deep dish pizza, taking my picture in front of the bean, figuring out public transportation, and did I mention deep dish pizza?!
My adventure here in Chicago is just beginning and I can’t wait!
This fall we had the wonderful opportunity to work with Cameo Chilcutt, a student of Northeastern Illinois University. Cameo spent the fall working in the lab and conducted her own growth chamber experiment with seeds from Selena’s summer REU project. Cameo was a great addition to the lab and asked some cool scientific questions about how water stress and maternal competitive environments affect germination in Lasthenia californica. Check out her final report posted below. We’ll miss having Cameo around but wish her luck in her future scientific endeavors!
Cameo Final Paper

Here at the Echinacea project, we like to count things. We like to count achenes, pollen, seeds from liatris and solidago, and really anything that will get us data. So, I am happy to give our (almost) full counting report from 2018.
This year, members of Team Echinacea counted over 717,000 achenes.
I, Personally, am wowed by this number,for not only does it mean that our team had to sit and click a computer screen 717,000 times, but it also means that we have roughly 1/3 that number of achenes sitting on the shelf right now! And that’s just from this year!
Thanks all for a great year of counting, and I hope we count even more next year!
Michael
Here at the Echinacea Project, we have some pretty rigorous protocols to make sure that everything goes right. This applies doubly so when it comes to ACE — the process we use to make sure we count all of the achenes on every head (hence the acronym, Accurately Count Everything). The process starts when we harvest the head in Minnesota in August or September, and ends sometimes years later when we have classified the last xray, with rigidly defined steps in between. This process leads us to a data set that has a human error rate of less than 1%
This is a story about that <1%
In fact, I would argue that this is a story about the 1% of that <1%. A head that has not only 1 terminal error associated with it, but 3.
This is the story of CG2016 XD-9460.
My experience with this head, whom we’ll nickname XD, began when Tracie gave me a list of the necessary rescans for 2016. These are the heads that, for some reason, never had their achenes scanned. They can be particularly troublesome because usually their achenes are in 3 separate places as part of ACE protocol that normally comes after the scanning step. I had gotten XD’s achenes from 2 of the 3 locations, and went to look for its orange coin envelope — the envelope that contains most of its achenes.
It wasn’t in the first place I looked, where it should be.
It wasn’t in the second place, where it shouldn’t be.
Nor was it in the third, fourth, or fifth places I looked, where it definitely shouldn’t be.
In fact, it was no where I looked. After an upsettingly long and painfully fruitless searching process, I decided to check and see who had cleaned XD, and maybe that would give me a clue where the envelope was. To my surprise, I saw the label that should have been on XD’s orange envelope still on the master label sheet, and the words “head not here” written in the notes section by the volunteer assigned to clean it.
How then, did we have some achenes for XD if the head was never cleaned? How did we have achenes if the head was never here?
This is an insight into our ACE protocol. Even with our organization and attention to detail, things can go wrong in ways we could never predict. After I completed my detective work, here is what I believe happened to CG2016 XD-9460. This is the verbatim note I have left in our “missing scans” log. It is unusual for this log to contain more than 10 words about each head.
1:56 12/5/2018 MCL
XD-9460 has more than one error associated with it. This is my best guess as to what happened.
Error 1. On the day it was harvested, the harvester came to a plant that had two heads on the ground: XD-9460 and XM-9011. The harvester put both heads in the same bag and made a note of it. I believe that bag was labelled XM-9011
Error 2. When it came to cleaning, XM-9011 was cleaned normally, and no note of a second head was ever made. I believe only one head was selected from the bag (either 9460 or 9011), labelled as XM-9011, and processed, while the other was ignored.
Error 3. During randomizing 9460 was not skipped, despite having no seed packet associated with it There is a white envelope and clear baggie with achenes “from XD-9460” in it; however, seeing as that is not possible, they come from a completely unknown source.
So there you have it. A story that took up a few hours of my time, so I thank you for taking a few minutes of your time to read it. If you’re wondering what happened to the data from this head, it has been labelled as “lost in the field” and wont poorly influence our analyses (or torture our lab managers) anymore.
Have a good weekend!
Michael
Hi flog,
Things have really gotten going, here at Echinacea headquarters. Julie, Sarah, Lea, and I have been spending much of our time cleaning and counting Solidago, and we’ve made pretty good headway, each counting about 2-3 samples per hour. We’ve also made some preliminary observations about variability. We’ve found high variability between plants, but not within plants! For example, we’ve seen ranges of 3 to 372 heads per stalk, and anything from 7 to 24 achenes per head in different plants. But within a plant, achenes per head is relatively uniform, generally ranging by ~2 achenes, which should allow Lea to precisely estimate total number of achenes per plant.
In addition to the Solidago, we’ve been working on Liatris. We finished the most important part of estimating potential reproductive success: x-raying the achenes to see if they contain seeds. This process involves placing baggies filled with Liatris achenes over a reusable x-ray film, shooting x-rays through the achenes, and quickly feeding the film through a machine that digitizes the film and allows for online counting of full and empty achenes. From this, we can estimate proportion of achenes containing seeds, multiply this proportion by number of achenes, and thus estimate reproductive potential of the plant. Because my project doesn’t involve seed set, Julie will be doing all the counting for Liatris (Fig. 1) while Sarah will handle counting for Solidago x-rays.
Until next time,
Tris

Fig. 1. Julie diligently counts full and empty Liatris achenes while Lea continues to count Solidago heads.
Hi Flog!
With half of our externship now past, some exciting things are happening in the lab! Yesterday, Sarah, Tris, and I had the opportunity to talk with Stuart, Michael, and Lea about some of the research questions and hypotheses we’ve started to form while working so closely with the specimens over the past week and a half. Stuart offered several suggestions about how to formulate these questions so that they are testable with the kinds of data we have, while also advising us about the statistical approaches we could take to assess our hypotheses. As we gather together the last crucial pieces of data, we’ll have the opportunity to try answering some of these questions. If all goes well, we will be rounding out our experience working with Team Echinacea by the end of next week with our own independent projects and analyses of the data.
In the meantime, we have been continuing our final steps of data collection. Despite our earlier trials and tribulations holding our breath while individually counting hundreds of seeds the size of sand grains, our new Solidago counting procedure seems to be a success! With a more efficient system in place, we hope to finish counting seeds by the start of next week. All that remains after that is x-raying and classifying the x-rays from all of our samples of Echinacea, Liatris, and Solidago before we can gather together all of our data into frames. As we learn more and more from our mentors about how to manipulate data sets in RStudio, I can’t wait to start assessing regressions and comparing models soon!
Excited for the number crunching,
Julie

Tris and Sarah demonstrating our new and improved Solidago counting procedure
Hi Flog!
As we near the halfway point of our externship with Team Echinacea, it’s time for a progress update! Julie, Tris and I have been knee-deep in data collection for the past week and a half: this includes both the “ACE” protocol with Echinacea achenes (cleaning, re-checking, scanning, counting, randomizing to create X-ray samples) and similar processing with Liatris and Solidago specimens. We quickly learned that the pappuses (papi? – the fluffy bits designed for wind dispersal) of Liatris and Solidago achenes adds a new level of difficulty to the counting and randomizing processes. Any air movement–including breaths–can cause major disruption to our work spaces, and the achenes themselves are often frustratingly small.
Though we finally made it through the Liatris samples by the end of last week, with this week commenced the Great Solidago Counting Problem of 2018. With minuscule achenes, of which there may be many hundreds on any given sample, Solidago is not easy to work with–especially when you’d like to estimate achene count per plant. We spent much of our afternoon workshopping several methods of estimation and randomization with Lea and Stuart, hoping to make the process as efficient as possible while also providing reasonably accurate achene counts for Lea to use in her ultimate data analysis. Our final method (still subject to change and optimization) consists of counting the number of heads on each sample, then randomly selecting five heads from which to count achenes. This way, we can extrapolate average achene count per head to the number of heads, then to the number of total florescences of the plant. We hope to power through the remainder of the Solidago samples within the next few days, then begin analysis of all the data we’ve collected!
Until then,
Sarah

Julie, happy to randomize some Echinacea rather than count Solidago
Hey, my name is Leah Rose and I am an intern from Lake Forest College! During the last few weeks, I have been working with Stuart Wagenius and Michael LaScaleia to research pollen limitation in Echinacea angustifolia to see if there is an effect on lifetime fitness when the flowers are loaded up with pollen and when they don’t get any pollen. I have loved getting a hands-on experience with researching these important native plants and helping make an impact on conservation of our native prairies! As a future high school biology teacher, experiencing science and conducting research is very important to me and I want to show my students that they can be a part of research such as this in the future. Science is for everyone and introduces you to people and subjects outside the usual scope of our lives!
Here is a picture of me totally beating Emma’s achene count! #winning

|
|