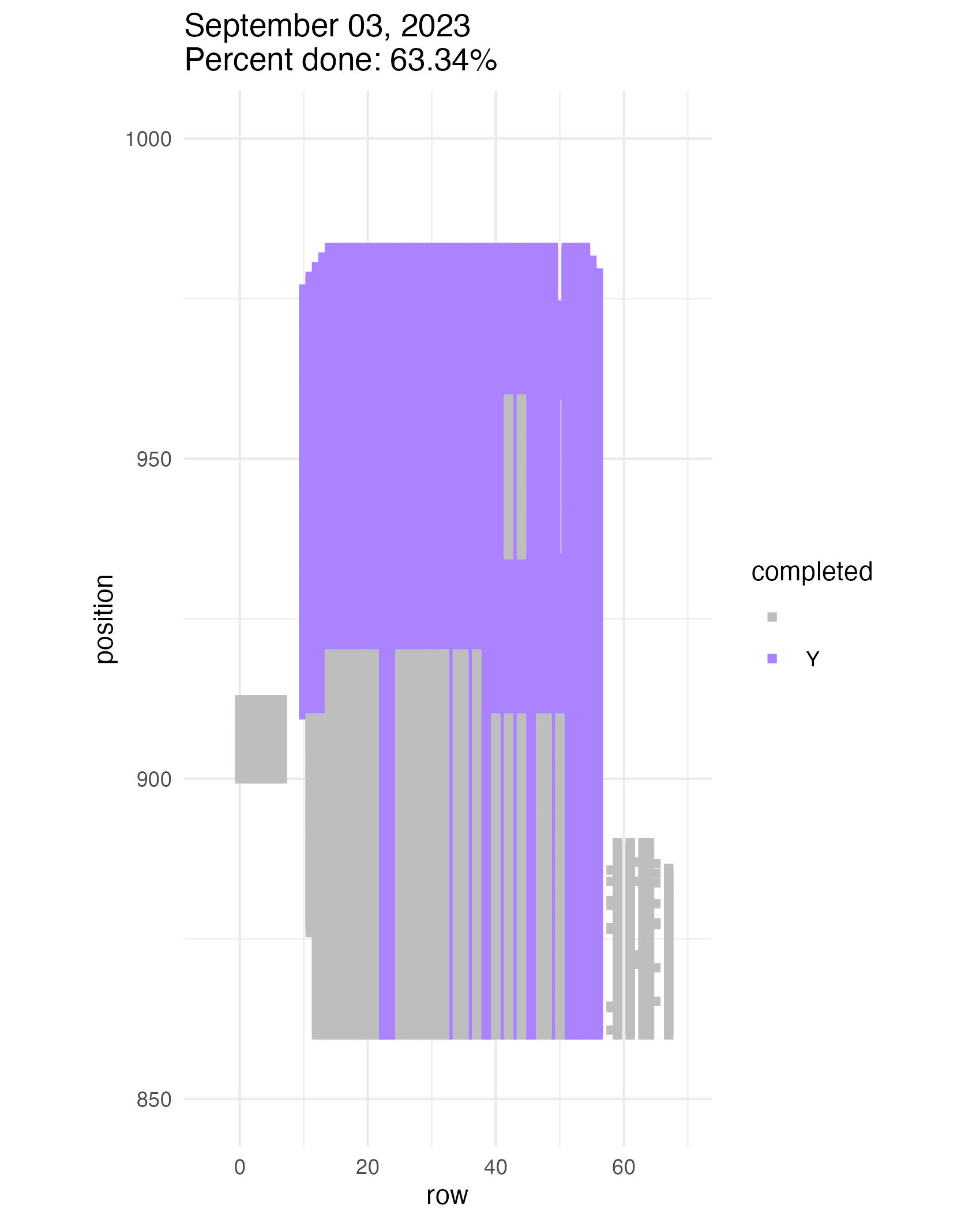
Let’s consult our total demometer to see how much progress we’ve made. The team has tackled some large sites in the past few weeks. We’ve made lots of progress! We’re about 45% done with all locs.
|
||||
|
The team has dwindled, and so has our progress on p1 this last month… With other priorities such as demo/surv and e-traps wrapping up in the next couple weeks, hopefully we can get back to it! 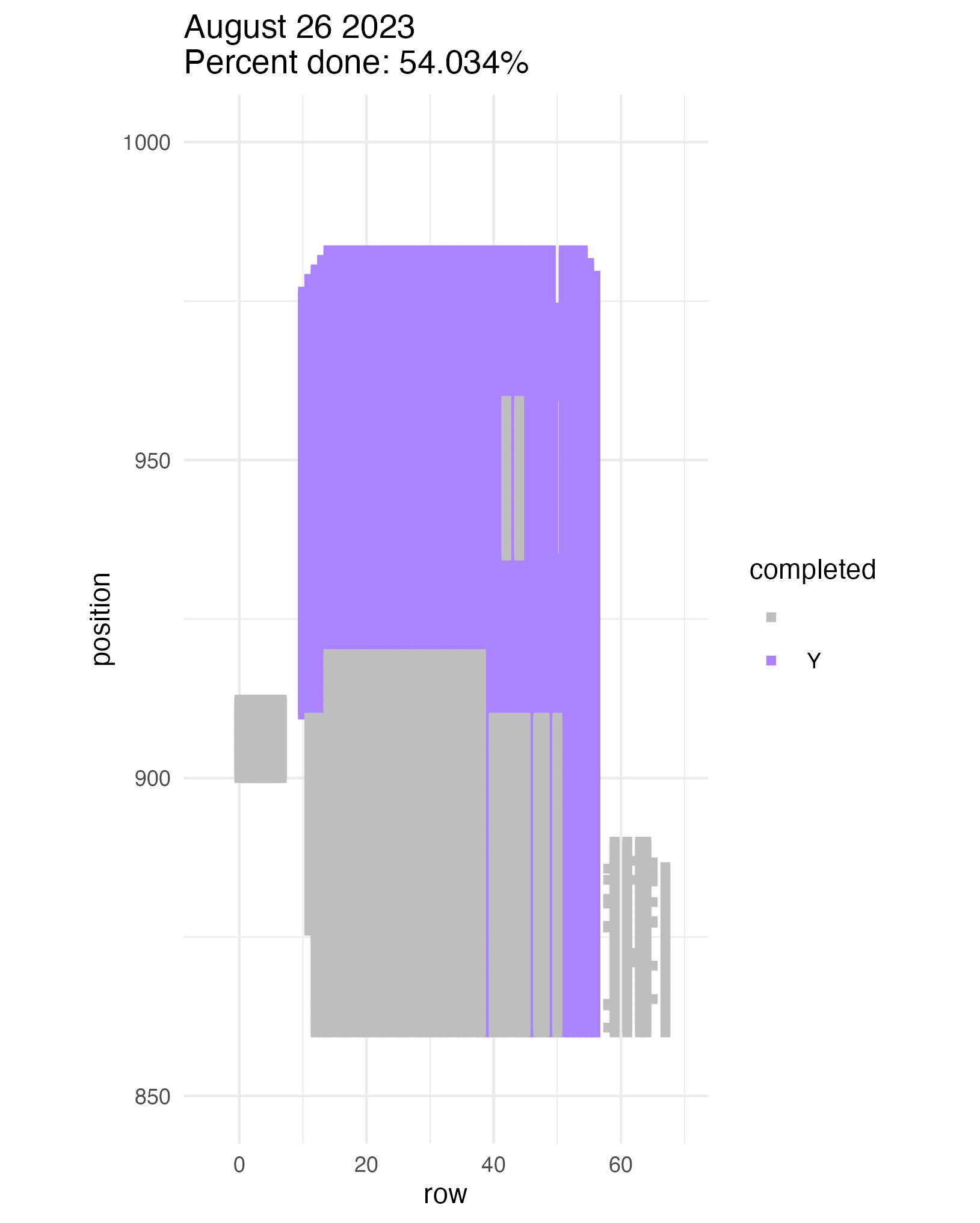 Members of Team Echinacea do a good job while measuring. Members of Team Echinacea are also humans (mostly, I think). We make mistakes! Maybe we searched for a plant that is there, but we couldn’t find it. Or maybe we had missteps while recording data. That’s why an essential part of our work each summer is identifying errors and inconsistencies in our database and then revisiting these in the field to check our work, as well as searching again for any plants we marked as can’t finds. Thanks to the ghost of interns past, there is quite a bit of framework and functions that already exist to handle this work. It’s also a bit of a hodgepodge to keep track of, especially when you don’t have last year’s cg intern to show you around (we miss you Lindsey!) Here, I am documenting my personal journey in figuring out the who what when where why and hows of this process. I’ll continue updating this as I learn more. Step 1: preRaw -> raw The first step in the process is to take the data from the visors, which we call “preRaw” and turn it into a “raw” state that we can more easily work with. This is mostly a quality control step that a) formats the data how we like it and b) identifies and eliminates any carriage return issues that would be a source of head scratching down the line. We do this step in a single script, but in separate steps for exPt08, exPt10, and then the rest of the plots as one. The same goes for the head data. Ideally, we would put in the work to combine all plots into one. Unfortunately there is a labor shortage, shall we say. 2024 script: ~cgData/summer2024/measureRaw/makeRawMeasure.R Step 2: raw -> good Next, we put the data through some more formatting tweaks. We also create a column called “recQ” (record quality) and tentatively mark every record we have as “good”. Any record with this field marked as “good” will be included in measureGood/plantAll_2024.txt and measureGood/headAll_2024.txt. Even prior to fixing errors, these can be useful in other processes, like setting up harvest. 2024 scripts:
Step 3: making “good” actually good That’s just the trick, isn’t it? There are a number of ways we go about getting good records. For some records, the mistake is obvious and easy to fix in R. When this is the case, you make the change in the offending column(s) directly in “pp” or “hh”, the plant and head measure data frames, respectively. If it’s as simple as updating plant status to basal, only the plant status field needs to be update and the recQ field remains set to good. If the entire record is bad, you can modify recQ to be something other than “good” to keep the record from being included in plantAll or headAll (e.g., “dup record”). Other times, we have to revisit the location in the field. When this is the case, we identify the issue and change recQ from “good” to a descriptive note about the issue. For example, any issue that I feel is worthy of a revisit in the field, I make sure to change the recQ to “revisit; xxx” with “xxx” being a brief note about the issue. The hh df is special in that it also has an “auditNote” column, where you can add additional notes about any issues. I’m not sure why this is exclusive to the head data. Some ways we identify issues:
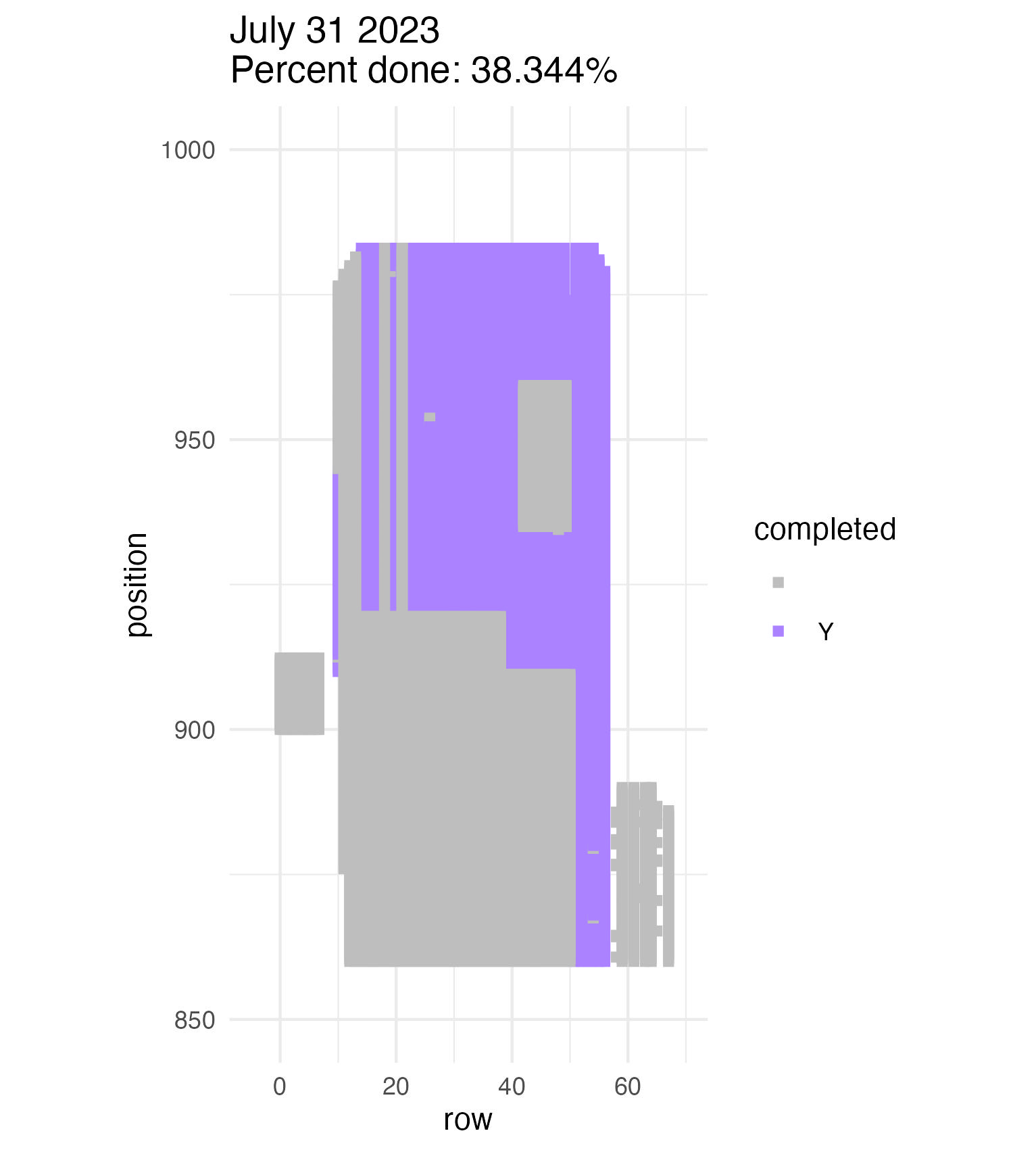
To be continued… We are busy with fieldwork here in western MN! Between measuring, floral assessments, emergence trapping, searching for flowering plants, and finishing up our pollen and nectar collections, we have our work cut out for us… We’ve been making steady progress with total demo. As of this morning, we have completed 34% of the locs where we search for echinacea. The team is making excellent time! These fast measuring skills no doubt transferred to speedy times at Flekkefest this weekend. 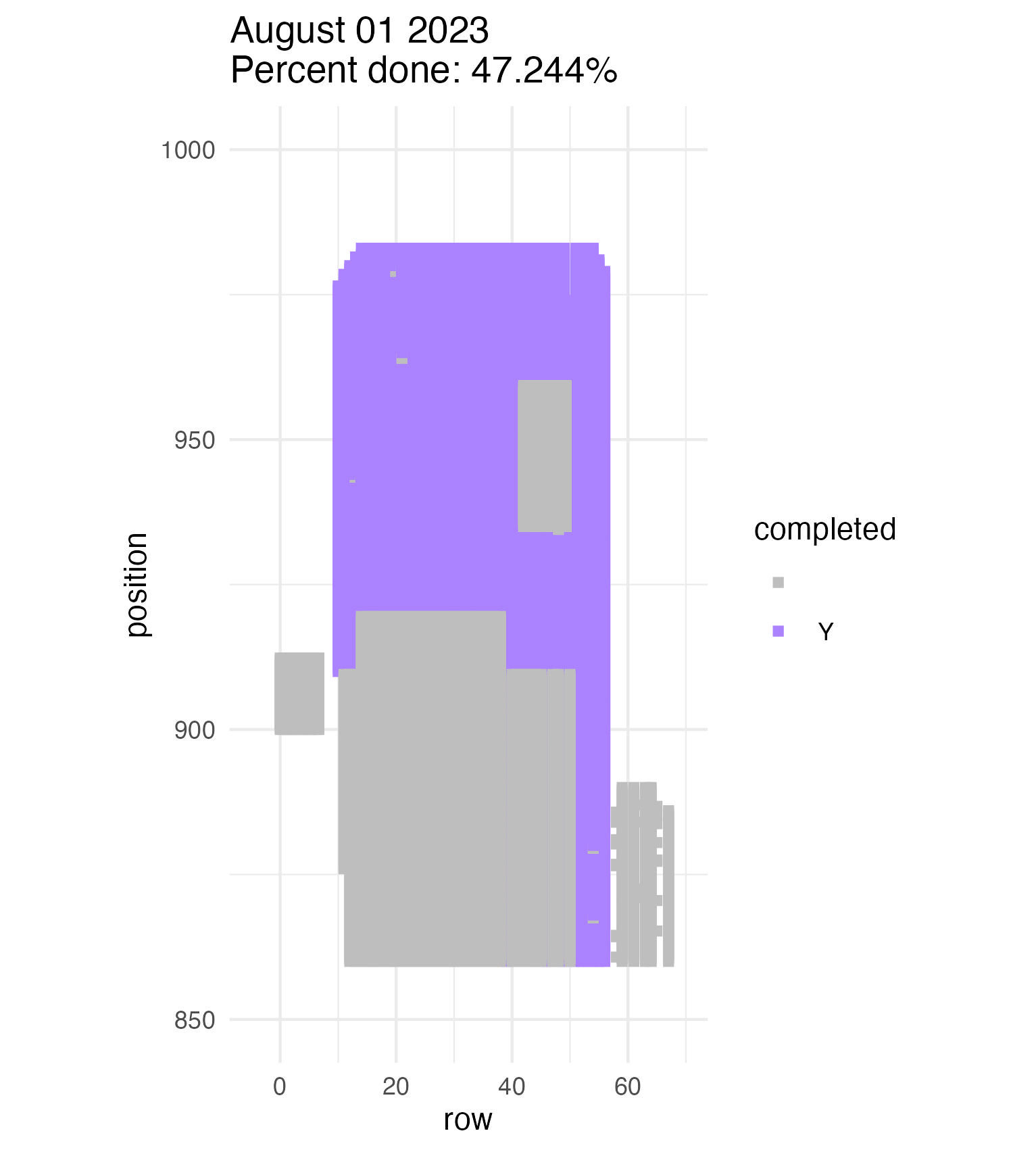 With a smaller crew and slightly soggy start to the morning, it was all-hands-on-deck as we continue to chip away at measuring experimental plot 1. We made great progress today, but still found time to stop and take pictures of cool critters! After lunch, we all headed out to retrieve and deploy emergence traps. It was a busy day for e-trapping, with three teams visiting eight sites in total.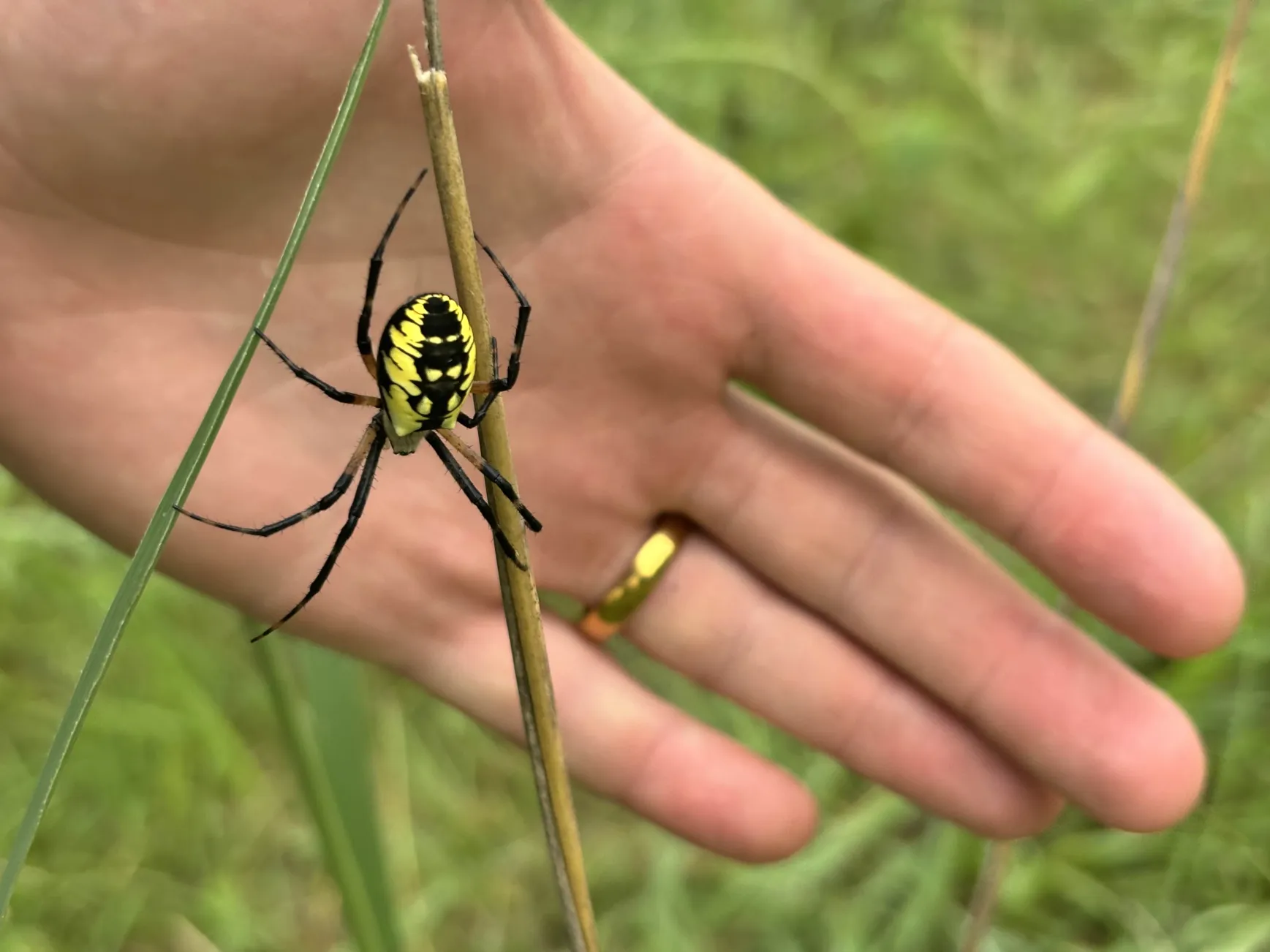   Small crews can make big strides! There were only 6 of us out there this morning, but we managed to crack 30% done.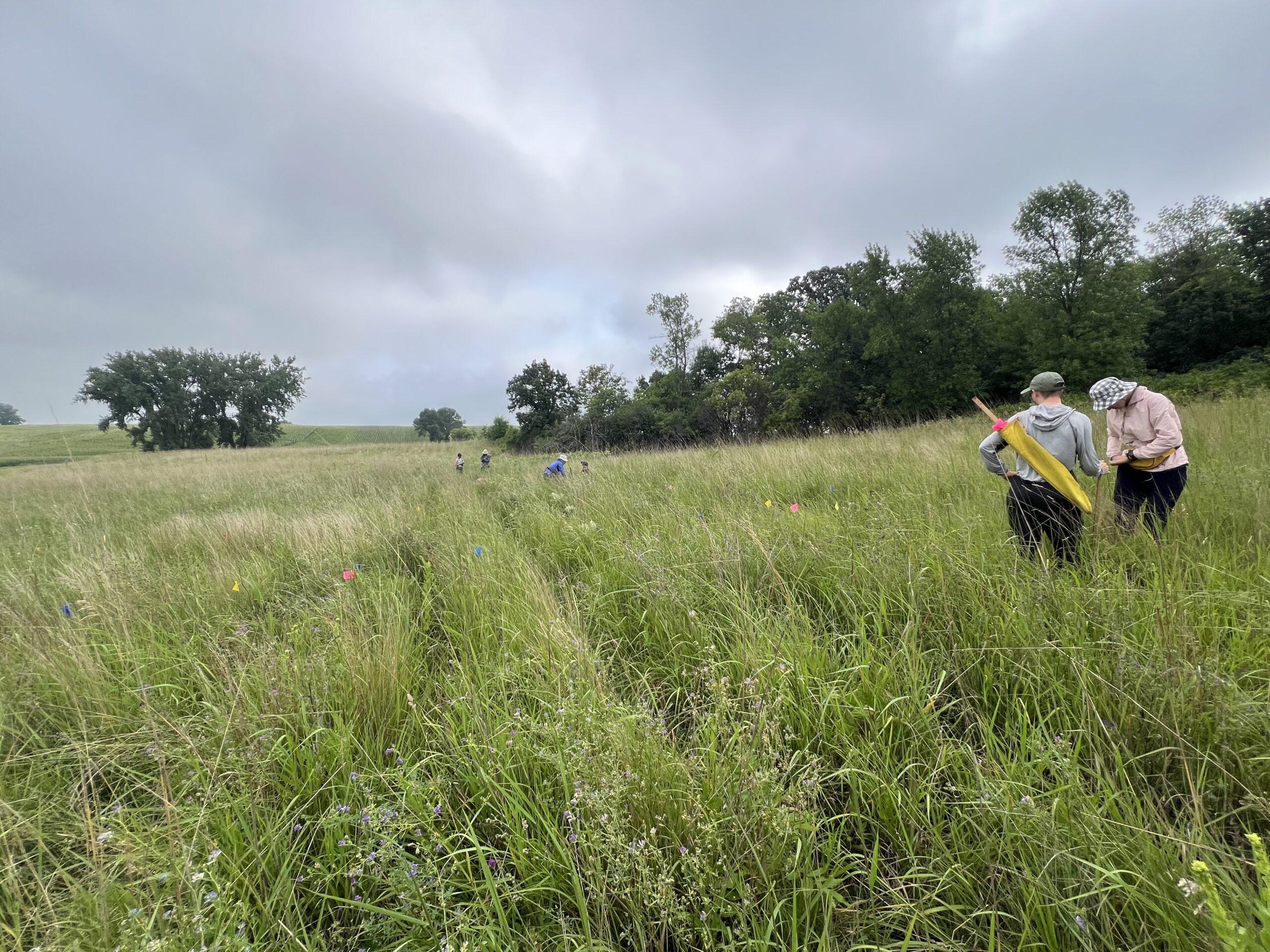 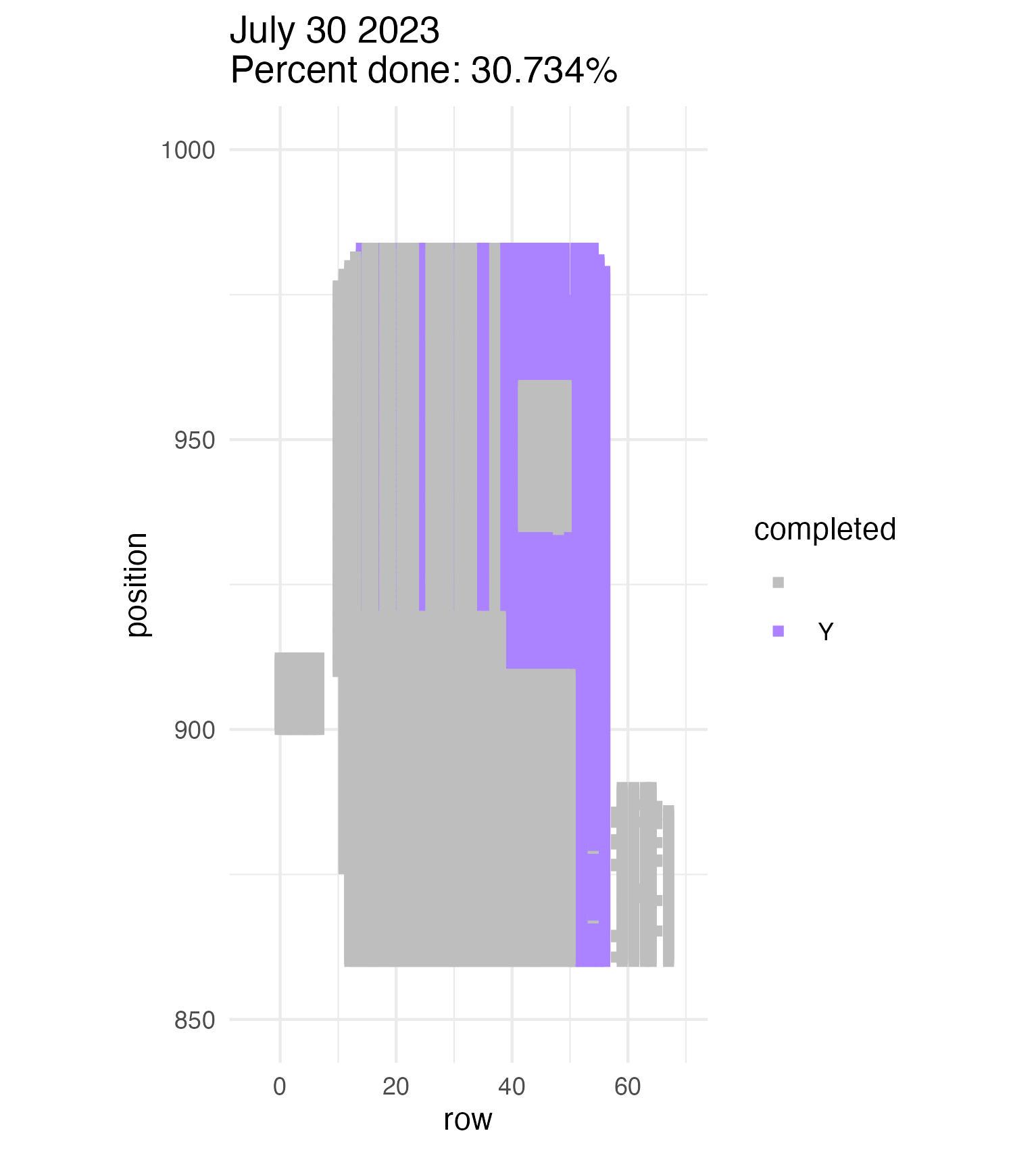  Friday, 26 July, 2024, was the first day we started measuring p1 this year! Coincidentally, this was the same day we started last year. This monumental task involves visiting 10992 positions in the plot and assessing the status of the plants there (or not, in the case of plants we have not been able to find in >3 years. RIP. Presumably.) After measuring again this morning, I booted up the old progress tracker (in the cgData repo, if you ever need it) to see how we’re doing. After making some adjustments (for the colorblind out there), here’s our progress over the last two days: 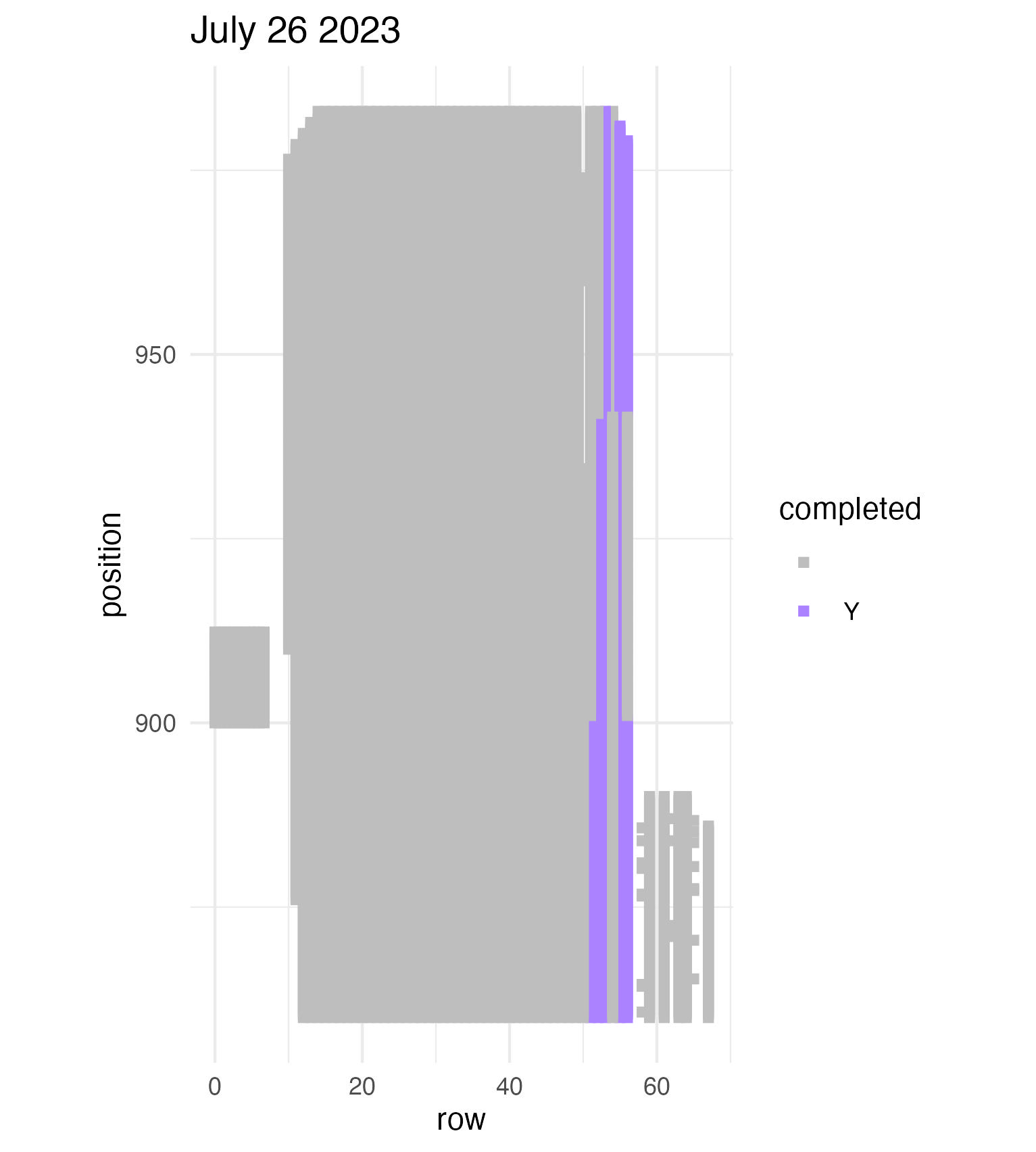 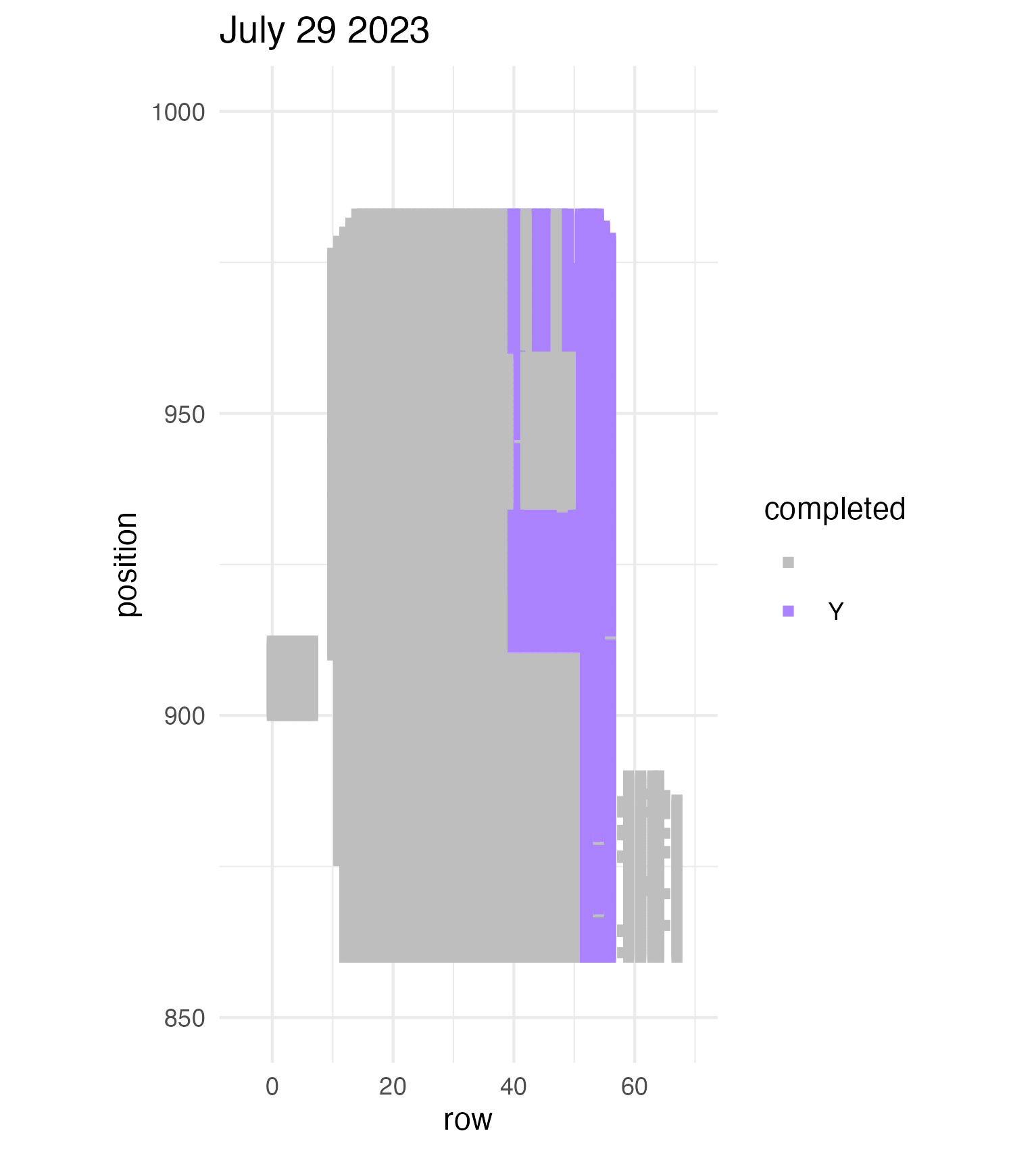 Today was also RET Brittany’s last day with the field team. Brittany, you will be sorely missed, and we hope you bring a bit of the prairie to share with your students!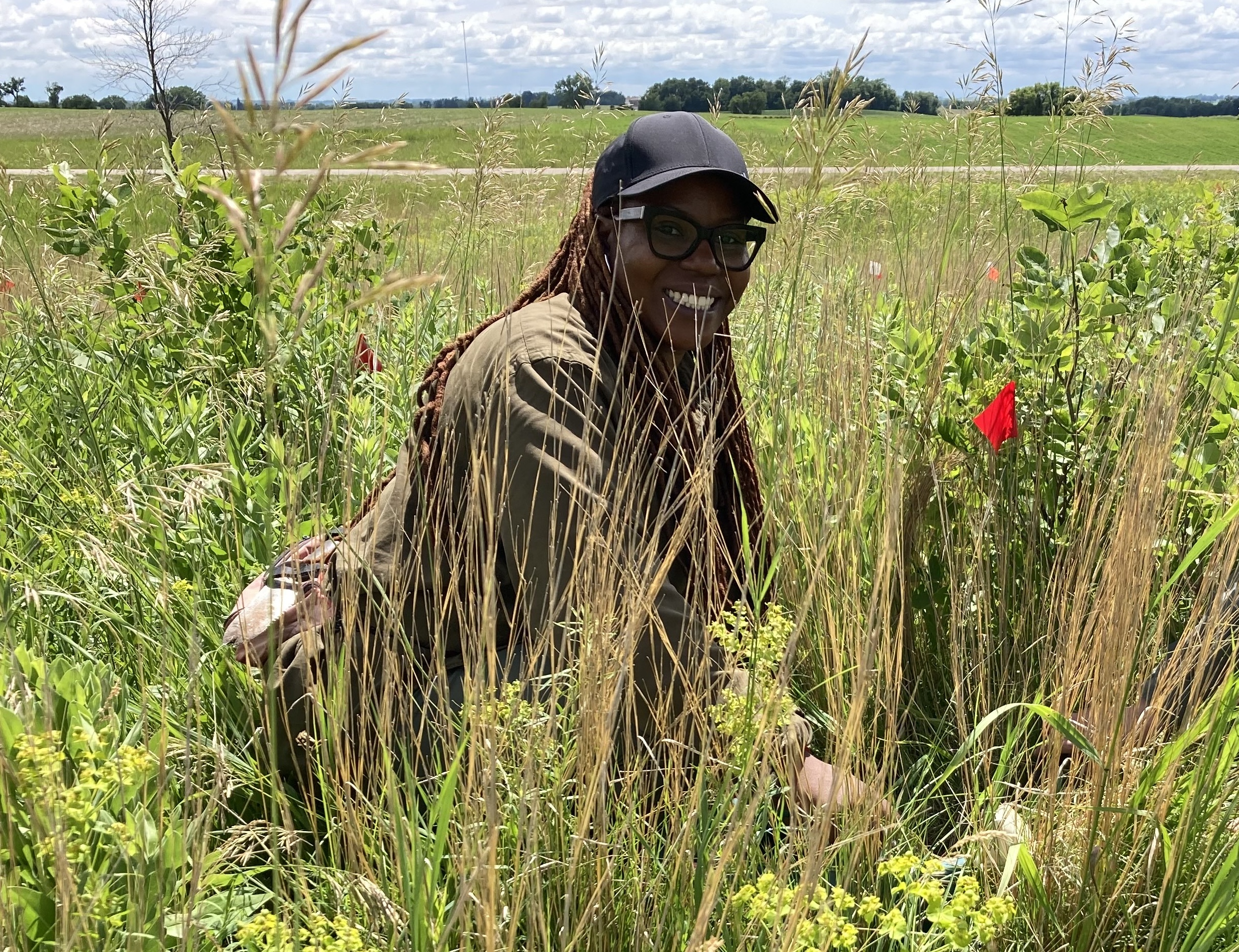  |
||||
|
© 2026 The Echinacea Project - All Rights Reserved - Log in Powered by WordPress & Atahualpa |
||||